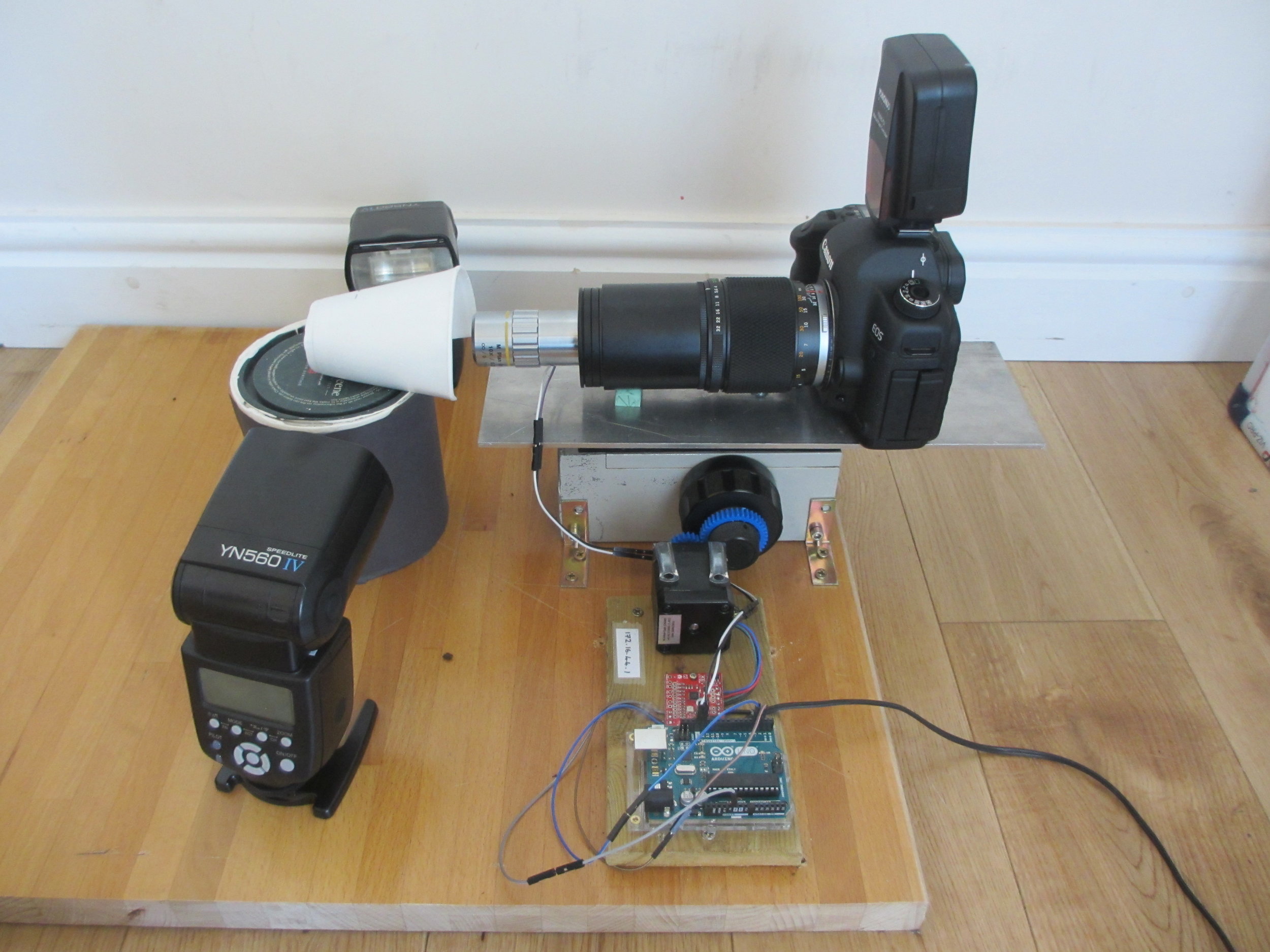
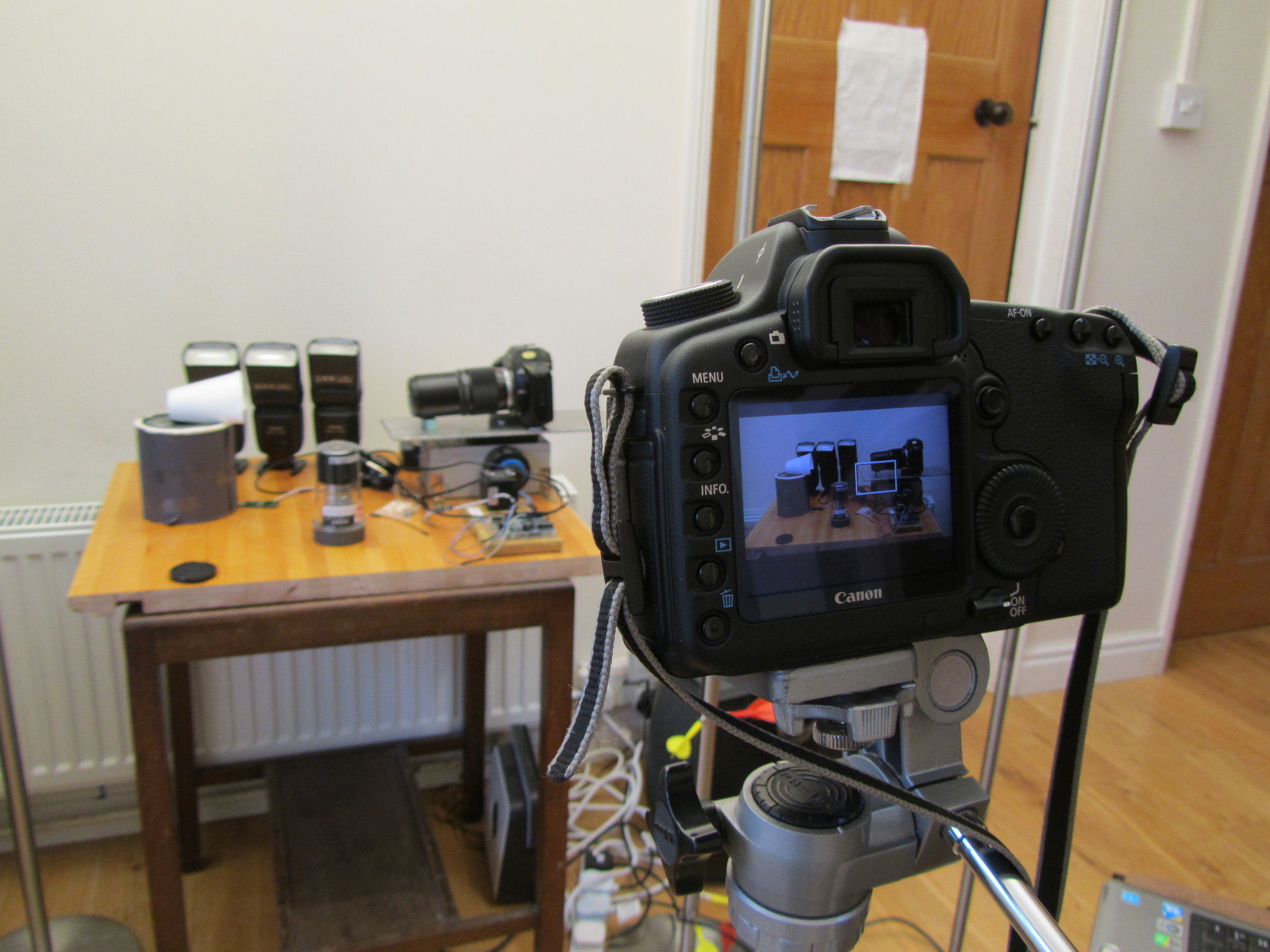
One of my funniest solved problems is that my DSLR is the only camera that I have that can record video, but it also has to appear in the videos. I got around this problem by putting my 27-year-old film SLR as a body double in the videos. The photo to the right shows my DSLR filming the focus stacking setup, with decoy camera body in place. It’s great fun editing the sound of the camera shutter into the finished video.
My other challenge is making these rather technical videos engaging to watch. There is a definite risk of them coming over as a bit dry, and so I try to keep them short and make the images interesting. I think that if I can improve my editing equipment at some point, I could make my videos much more engaging.
I’m really enjoying making educational videos and would like to keep doing this work after the end of the OpenPlant grant. I’ve been in touch with the University Public Engagement Office, who have been very helpful, and I’m hoping to learn some tips from them.
You have been awarded both a Biomaker Challenge and OpenPlant Fund grant. How have these enabled the development of the project?
JD: My work absolutely could not have been done without these grants. Most of the work has been done through collaboration, volunteer labour, and home engineering. However, the grants paid for the microscope objectives. Without these amazing lenses, I could not have done the work.
How do you feel the project is progressing?
JD: I think it's going very well. I have four good videos already online, and a lot of written documentation. I have registered a new domain (www.chlorophyllosophy.uk) as a central doorway to all of the material, and I still have lots of ideas for other videos to make.
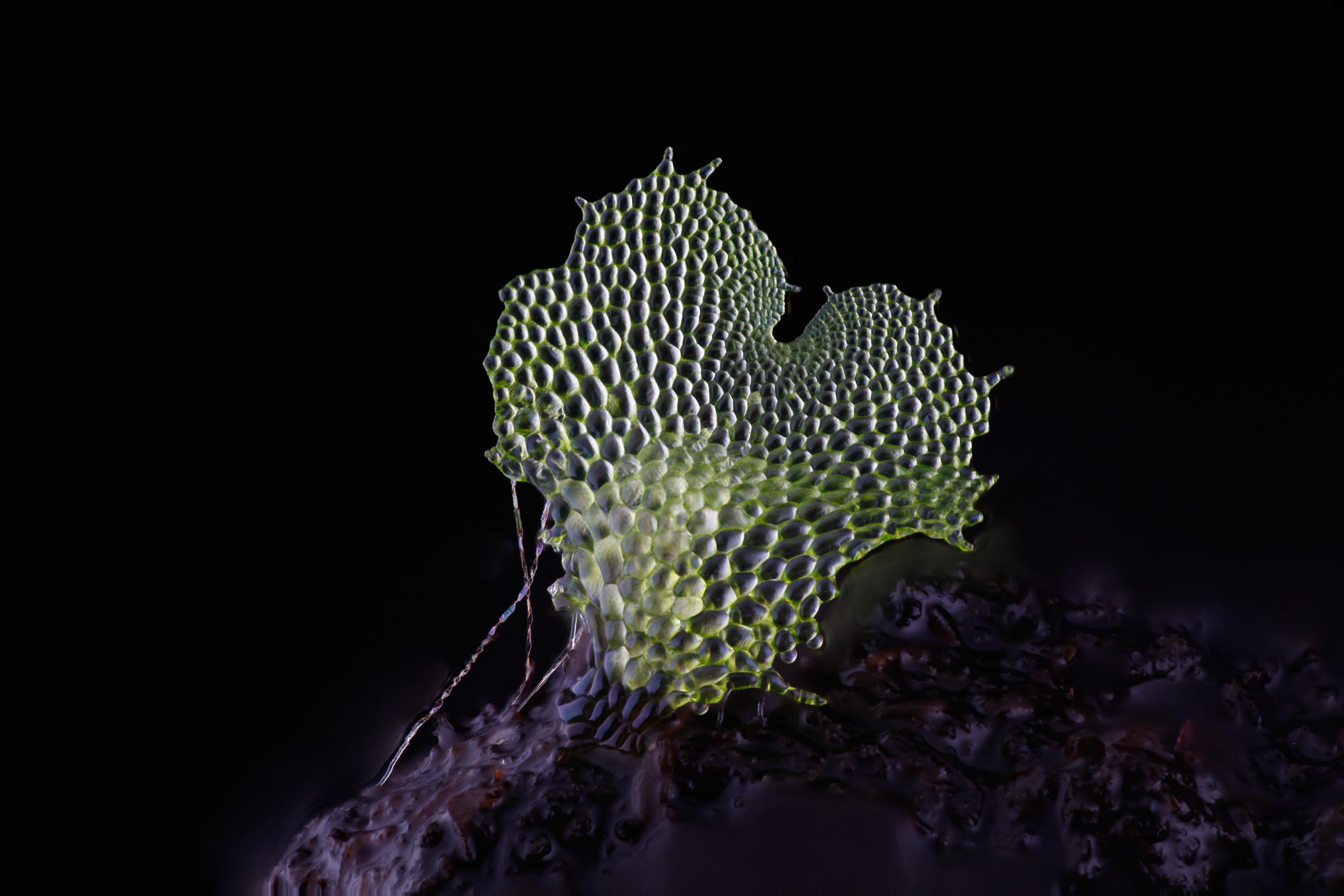
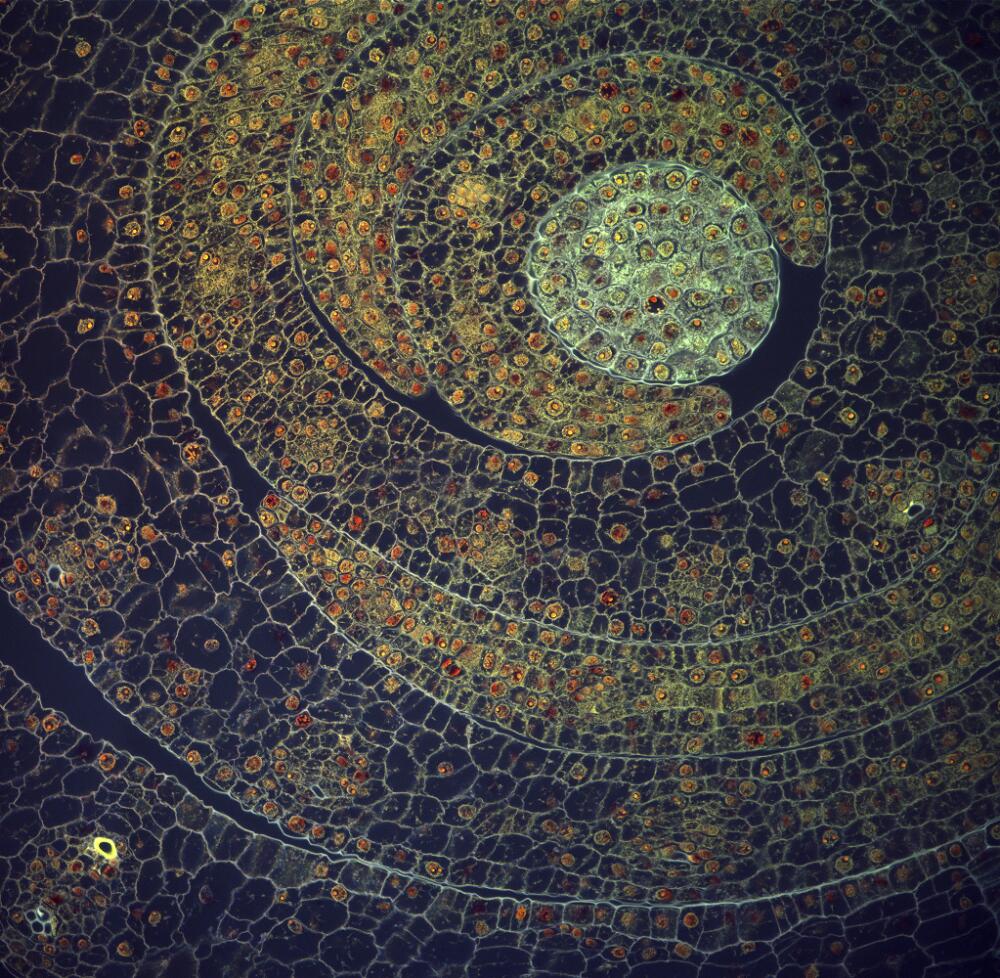
Two out of three of my lenses have arrived and I am looking forward to taking some great photos. My Utricularia gibba (bladderwort) plants are growing well in their casserole dish. Utricularia gibba is a small, carnivorous aquatic plant that develops traps to capture its prey. They are being studied by my collaborator Christopher Whitewoods at the John Innes Centre and I have already taken my first few photos of them, as the new traps develop. The traps have a beautiful structure, and as an aquatic plant, will be a great challenge to photograph.
I hope soon also to visit the Sainsbury Laboratory in Cambridge to photograph the trichome mutant phenotypes in Arabidopsis thaliana, belonging to my collaborator Aleksandr Gavrin. I really look forward to the challenge of photographing trichomes, that will have other trichomes behind to confuse my software.
I have also just sewn a new batch of fern spores and those plants will be a real treat to photograph when the time comes.
What are the future opportunities to take this project forward?
JD: One of the biggest pitfalls for photographers is that they become so fascinated by the stream of newer and better camera equipment, that they forget to actually take any photos. I think that in the next couple of years, it's very important that I actually take the time to take some photographs. With this new technology that I have built, and with the opportunity of my volunteer labour, these will add hugely to the body of research knowledge.
Jennifer's project is also documented on Github: https://github.com/BioMakers/Gametophyte-Fern-photography-2018/blob/master/README.md

![[Closes 24 Nov 2107] Apply now to the OpenPlant Fund!](https://images.squarespace-cdn.com/content/v1/54a6bdb7e4b08424e69c93a1/1509564315902-TUO4I6QRWI9TT8UGSIAJ/OpenPlantTwitter_400x400+%281%29.jpg)

![[Closes 7 Mar 2017] OpenPlant Research Associate (Haseloff Lab)](https://images.squarespace-cdn.com/content/v1/54a6bdb7e4b08424e69c93a1/1486552818859-FH76MCA8SMFU93WB85RX/OpenPlantTwitter_400x400.jpg)